iVici 0.9
For Mac OSX, Linux and Windows
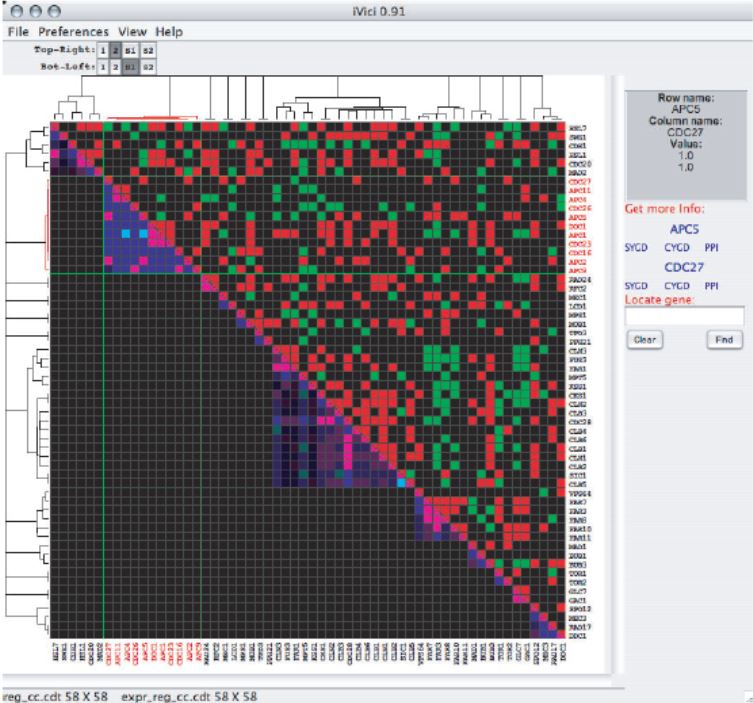 iVici is a Java application for analysis of cellular networks represented as addressable symmetric or asymmetric two-dimensional matrices. iVici is designed for simultaneous visualization and correlation of multiple datas sets, representing any relationship between a set of genes, mRNA or proteins. Visual overlay of datasets and addressable access to gene annotations allows for comparison of networks of different types, for example, protein-protein interactions and genetic networks or investigating the dynamic reorganization of a particular network.
iVici is a Java application for analysis of cellular networks represented as addressable symmetric or asymmetric two-dimensional matrices. iVici is designed for simultaneous visualization and correlation of multiple datas sets, representing any relationship between a set of genes, mRNA or proteins. Visual overlay of datasets and addressable access to gene annotations allows for comparison of networks of different types, for example, protein-protein interactions and genetic networks or investigating the dynamic reorganization of a particular network.
iVici is a multi-platform application written in Java. Installers for Mac OS X, Linux and Windows are available for download.
For more information on iVici, please refer to the online documentation.
MotifHound 1.0
For Mac OSX, Linux and Windows
MotifHound is an algorithm for the discovery of degenerate linear motifs encoded in protein sequences. MotifHound is based on the fundamental premise that, linear motifs mediating a particular function are enriched in proteins exhibiting that function and rare or absent in other proteins. Accordingly, MotifHound exhaustively enumerates all possible motifs present in proteins of interest, including all degenerated forms, and scores the biological relevance of each motif by evaluating its enrichment relative to the proteome using the cumulative hypergeometric distribution. The package includes also a benchmark of 11,880 protein sets from S. cerevisiae in which we artificially spiked-in motifs varying in terms of three key parameters, (i) occurrences, (ii) length, (iii) and number of degenerate positions. The benchmark enabled to evaluate the impact of these three properties on the performance of MotifHound.
MotifHound package is available for download and online at OMICtools.
For more information on MotifHound, please refer to the online documentation.
DALEL 1.0
Webserver for internet users
DALEL algorithm exhaustively searches the linear information in groups of related proteins. First, it enumerates all possible motifs of variable length including any number and combination of wildcards. Then, exhaustively explores all possible combinations of amino acids at distinct positions, to discover residues with preferences for individual or multiple amino acids. The use of parallel suffix tree data structure allowed the exhaustive search of linear motifs; an approach that has previously been physically unfeasible because the combinatorial space of sequences is too vast. DALEL comprises also a statistical method for inferring binding specificity of linear motifs to individual recognition modular protein domains. DALEL webserver is available at http://michnick.bcm.umontreal.ca/dalel/.
VertexSort 0.1-1
For Mac OSX, Linux and Windows
VertexSort is an R package that permits to apply the Vertex Sort algorithm (Jothi et al. (2009) DOI: 10.1038/msb.2009.52) to a graph in order to elucidate its hierarchical structure. It also allows graphic visualization of the sorted graph by exporting the results to a cytoscape friendly format. Moreover, it offers five different algorithms of graph randomization: 1) Randomize a graph with preserving node degrees, 2) with preserving similar node degrees, 3) without preserving node degrees, 4) with preserving node in-degrees and 5) with preserving node out-degrees VertexSort is an R package that permits the application of the Vertex Sort algorithm on an interaction network.
VertexSort is a multi-platform package written in the R scripting language and is available for download.
For more information on VertexSort, please refer to the online documentation.
Plasmid requests
The Michnick Lab has deposited plasmids at Addgene for distribution to the research community. Addgene is a non-profit plasmid repository dedicated to improving life science research.
Plasmids are available at http://www.addgene.org/Stephen_Michnick/.
At the moment, only plasmids to perform IFP-PCA in yeast and mammalian cells are available (Tchekanda et al. 2014, Nat Methods). Our Addgene plasmid library will be updated soon.